Main Content
Researching the principles that control gene regulatory circuits
Investigation of how genes in yeast and mammalian cells are wired up and which mechanisms control these gene networks will help understand cell differentiation. For this, we combine experimental methods with mathematical modeling.
Genetic networks often rely on feedback control, chromosomal epigenetic process and regulation of RNA turnover to promote adaptation, cell decision and stochastic differentiation.
Feedback regulation of adaptation and cellular memory
We develop mathematical models to assess how feedback regulation can improve cellular adaptation and memory; we also test these predictions experimentally.
Measurement and estimation of in vivo systems parameters
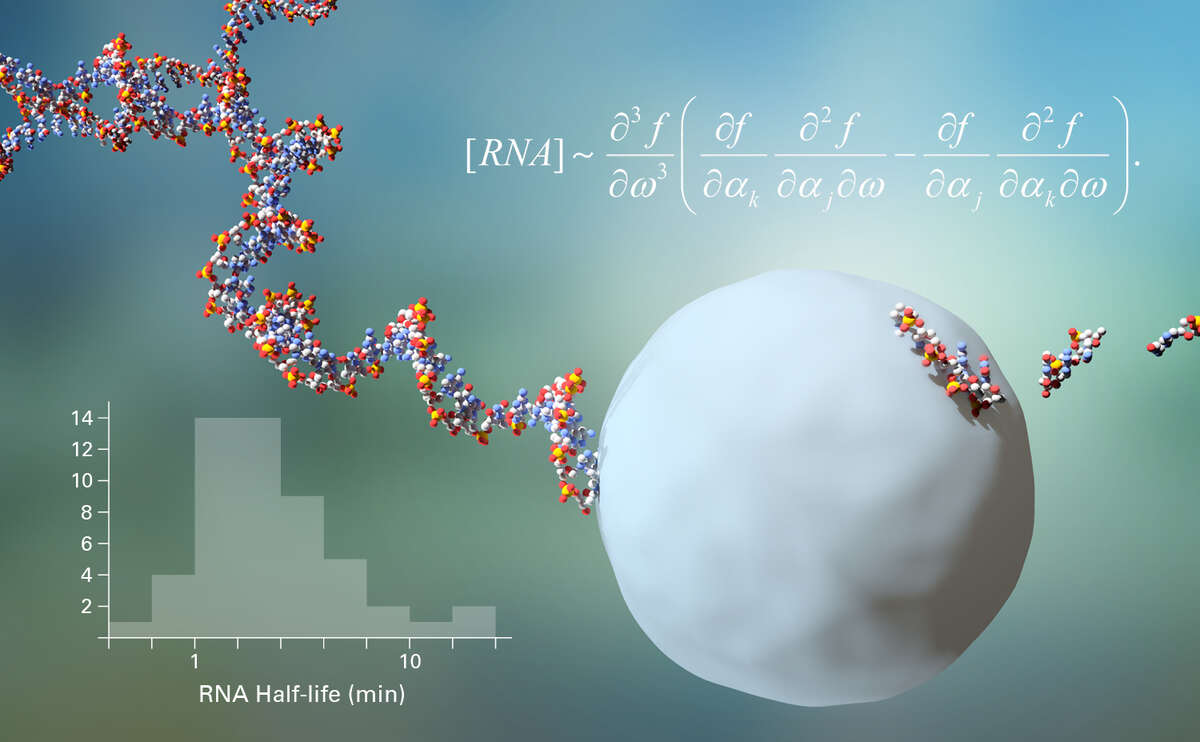
Adaptation depends on RNA and protein turnover. We develop approaches to measure these parameters and we are exploring how to find a consistent set of parameters, when different methods to measure RNA half-lives yield different results, by estimating the underlying true parameters.
Stochastic and epigenetic processes in cellular differentiation
Genes in higher eukaryotes are regulated by long-range interactions, which can determine if and what combinations of genes are expressed in a chromosomal segment. Many of these long-range interactions are mediated by epigenetic modifications of the genes. We found that epigenetic silencing in yeast can support cell fate decisions only when both ends of a gene are silenced. We also study stochastic gene choice during differentiation of embryonic stem cells to neurons. We explore how the combinatorial control of epigenetic signatures can help to diversify neuronal phenotypes.