Main Content
We study the cellular organization and regulation of metabolism by large macromolecular assemblies as well as giant microbial biosynthetic assembly lines. Our research builds on hybrid use of state-of-the-art cryo electron microscopy, X-ray crystallography and advanced optical microscopy to elucidate blueprints of dynamic protein and principles of their assembly and cellular level interplay.
Cellular Organization and Regulation of Metabolism
Metabolism in eukaryotic cells is tightly regulated in response to nutrient and energy availability as well as growth factor signaling for coordination of metabolism across organs and organism. Metabolic reactions are spatially organized not only into organelles but also by the transient formation of localized protein assemblies in the cytosol. Two key focus areas of our research with outstanding relevance for human health are the mTOR kinase, the central regulator of cellular growth and proliferation, and fatty acid and lipid biosynthesis.
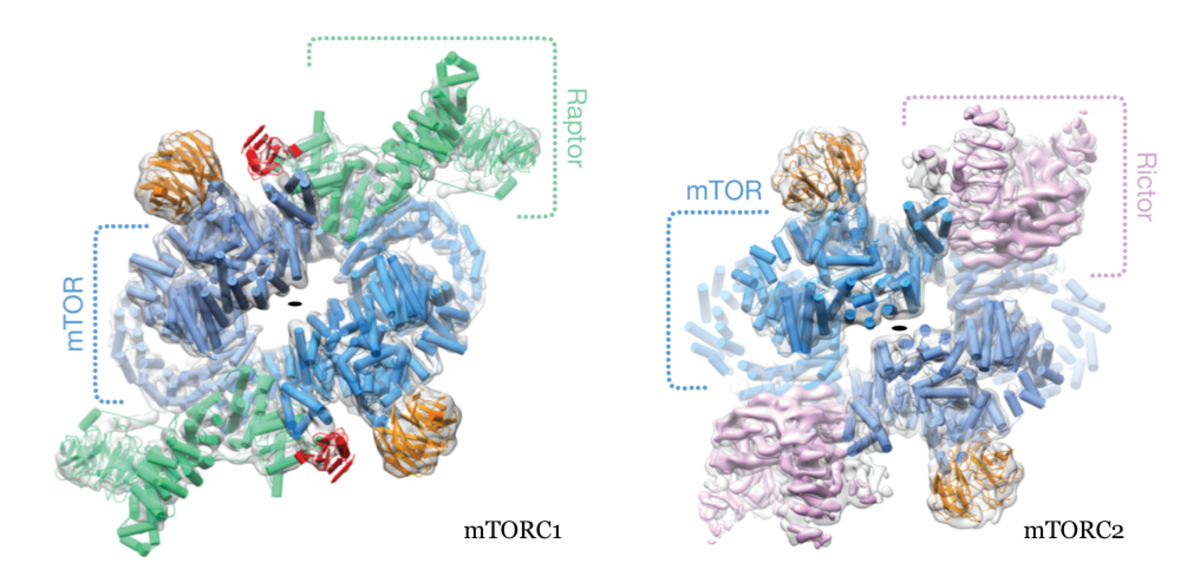
The mTOR kinase exerts its functions in two functionally and structurally distinct complexes, mTORC1 & 2. The mTORCs integrate multiple input signals from nutrient levels, cellular energy state and external growth factors for the control of anabolic vs. catabolic metabolism, proliferation and growth as well as cytoskeletal dynamics and cell survival. Major downstream targets include fatty acid and lipid metabolism, protein and nucleotide biosynthesis. Due to their central role in driving cell proliferation, mTORCs are key targets for anti-cancer therapy.
The two complexes share the defining core mTOR protein but employ characteristic subunits, Raptor for mTORC1 and Rictor for mTORC2, for capturing input signals controlling and localization, as well as for recognizing specific targets for phosphorylation. We have resolved the architecture of human mTORC1and mTORC2 (Fig. 1) by a combination of crystallographic studies and cryo electron microscopy in collaboration with Mike Hall (Biozentrum) and Nenad Ban (ETH Zurich). Current work focusses on high-resolution structural studies and analysis of transient interaction dynamics to reveal principles of substrate recognition, regulation and localization.
Lipids are an important source of natural chemical diversity and contribute to signaling, trafficking and inflammation. Lipid compositions reflect the metabolic and functional state of a cell. Impaired lipid and fatty acid metabolism plays a considerable role in the pathogenesis of major threats to human health, including type 2 diabetes, fatty liver disease, atherosclerosis and cancer. A central aim of our work is to improve our understanding of the regulation of fatty acid and lipid metabolism. Eukaryotic lipid and fatty acid metabolism remains a critical challenge for studies at the atomic, molecular and cellular scale: In contrast to simple prokaryotic systems, fatty acid metabolism in eukaryotes builds upon large multienzymes, which integrate multiple catalytic activities into giant proteins. Later steps of lipid metabolism are occurring in membrane space and rely on membrane-integral or -associated proteins. Combining experimental structural studies with functional analysis, we elucidate fundamental principles of multienzyme architecture & regulation and the role of protein assemblies for spatial organization of metabolism.
A landmark example are giant eukaryotic fatty acid synthases (FASs), which comprise seven types of functional domains and carry out more than 40 reaction steps for the biosynthesis of fatty acids from acetyl- and malonyl-CoA. Our structural studies have revealed the intricate interlinking, coupling and crosstalk between domains that result in emergent properties beyond the functionality of their isolated building blocks and shed light on the evolution of this family of multienzymes (Maier et al., Science 2008; Maier et al., Quart Rev. Biophys. 2010; Bukhari et al., Structure 2012; Herbst et al., Nature 2016)
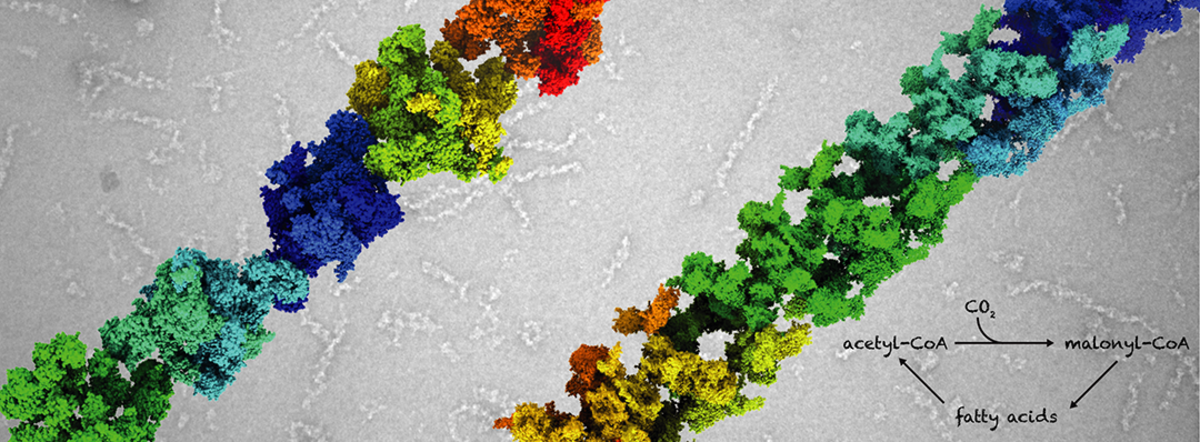
Acetyl-CoA carboxylase (ACC) catalyzes the committed and rate-limiting step in fatty acid biosynthesis, the two-step ATP-dependent carboxylation of acetyl-CoA to malonyl-CoA. It serves as a hub for short-term regulation by allostery, post-translational modifications and direct protein-protein interactions. Since more than 50 years it has been known that the control of human ACC activity is linked to polymerization of ACC into giant filaments. However, the link between polymerization and activity remained enigmatic. Our recent structural studies have revealed the architectures of fungal and human ACC and uncovered how the multienzyme architecture supports unique mechanisms for the consistent regulation fatty acid biosynthesis. In particular, our cryoEM analysis of ACC filaments revealed that filament formation and direct protein-protein interactions exert control of enzyme activity by locking ACC in distinct active or inactive conformations (Fig. 2).
We currently aim to fully reveal the multitude of regulatory options for human ACC as well as the role of multienzyme filaments for spatially organizing and localizing anabolic metabolism in the cytosol.
Microbial Assembly Lines for Polyketide Biosynthesis
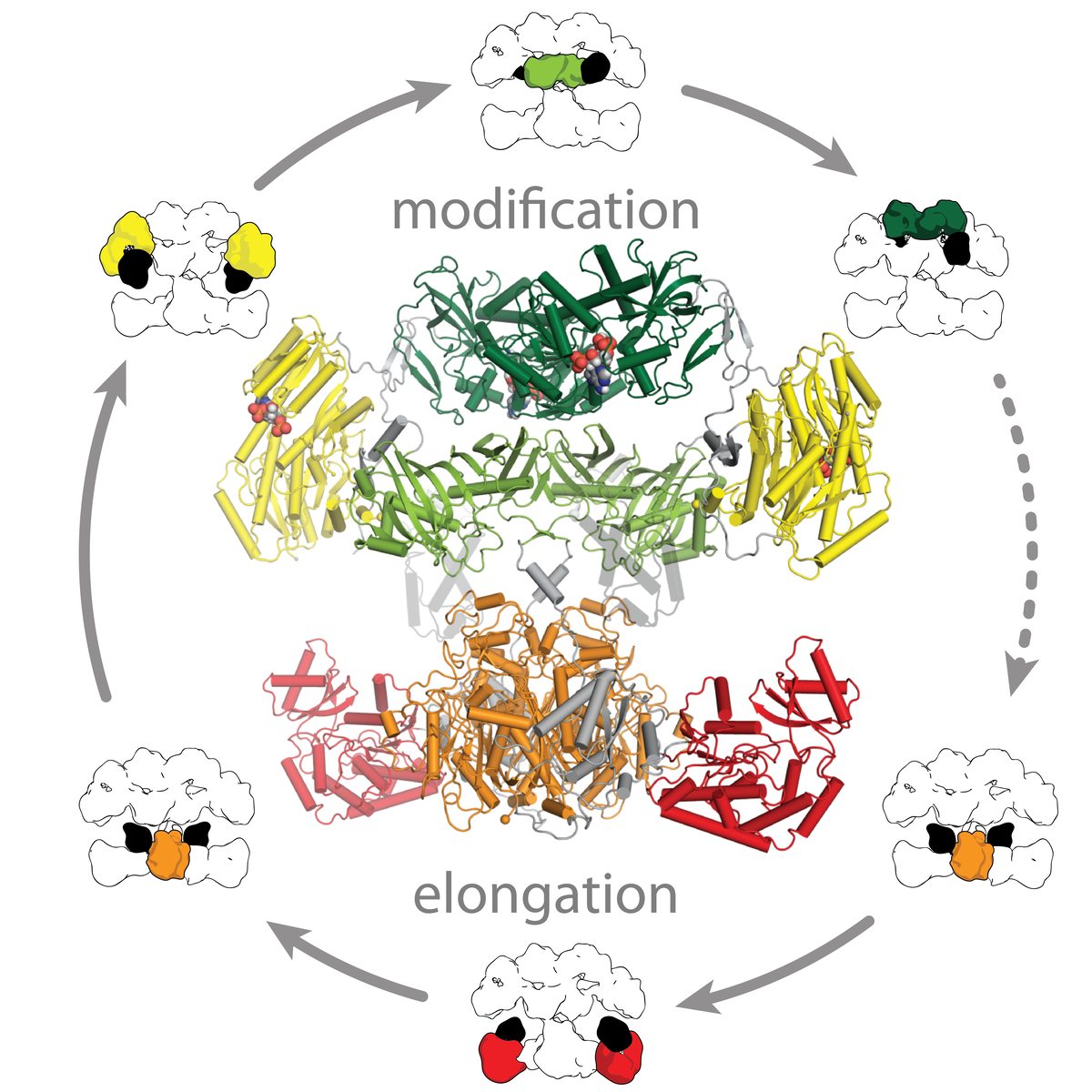
Polyketide synthases (PKSs) are microbial assembly line proteins and arguably the most complex and versatile biosynthetic multienzymes. Their highly diverse and complex polyketide products include compounds with outstanding biological activities, e.g. antibiotics and immunosuppressants. Polyketides are amongst the promising sources of novel drug candidates.
Polyketide biosynthesis is achieved either by multiple cycles of iterative action of one PKS module or by a directed handover of substrates between modules. PKSs modules share a common dimeric architecture based on a structural and functional segregation into two multidomain polypeptide regions: The condensing region is responsible for precursor elongation, the modifying region for processing of intermediates. PKS employ internal carrier protein domains (ACPs) for shuttling of covalently tethered intermediates.