Main Content
Chromosomes in 3D
The developmentally dynamic nucleus
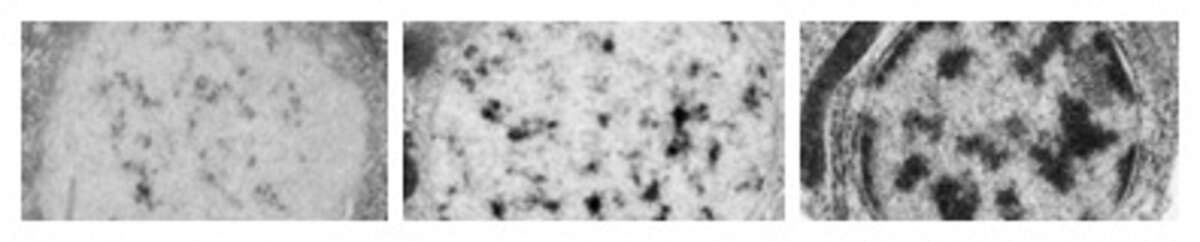
How are chromosomes organized within nuclei? And how does organization impact developmental events? The nucleus is very dynamic during embryogenesis, in part because heterochromatin is made de novo, as shown below for young, pre-gastrula embryos. Our goal is to understand how the nucleus establishes its mature structure, what factors regulate the process and why it matters to animals. Recently we discovered a complex that times the onset of heterochromatin, which you can read about here * We also developed a new method called Chromosome Tracing to determine the conformation of chromosomes at the single molecule level (below). These and other projects are giving us an unprecedented view of nuclear organization during embryogenesis, and revealing how the early embryo is drastically different from the differentiated state.
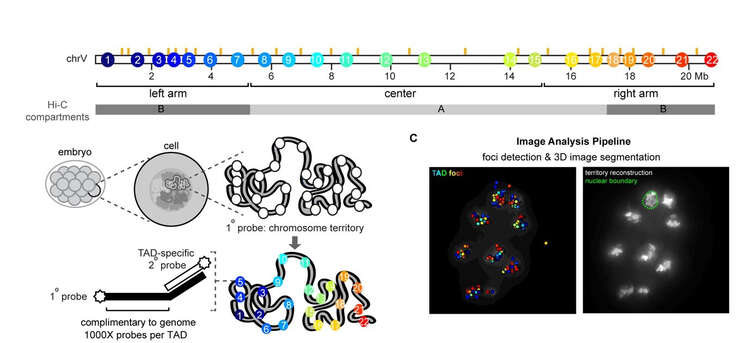
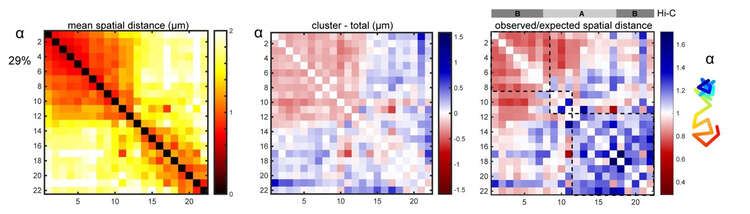
FISH probes are designed against a region of interest, and rounds of FISH imaging allow us to track the conformation of chromosomes in intact animals. We can calculate the folding properties and also examine mutants to see how they impact normal folding. From these sorts of analyses we’ve learned that worm chromosomes look like a barbell, with a lot of folding in the ‘arm’ regions and less in the center. Below is an example of what a wild-type chromosome looks like. We now will apply this method to a range of biological questions such as how do these structures change during development.
Read more:
Sawh et al., Lamina-Dependent Stretching and Unconventional Chromosome Compartments in Early C. elegans Embryos. Mol. Cell, 2020.
Mutlu et al. MET-2, a SETDB1 family methyltransferase, coordinates embryo events through distinct histone H3 methylation states. Development, 2019.
* Mutlu et al., Regulated nuclear accumulation of a histone methyltransferase times the onset of heterochromatin formation in C. elegans embryos. Science Advances, 2018.
What got us started:
Yuzyuk et al. The Polycomb Complex Protein mes-2/E(z)Promotes the Transition from Developmental Plasticity to Differentiation in C. elegans Embryos. Dev Cell, 2009.
Photo: Frederick Blichert dgit.com, David Hall, Ahilya Sawh